Data and Resources
TP53 mutational status is predictive of pazopanib response in advanced sarcomas.
K. Koehler,1 D. Liebner,1,2 and J. L. Chen1,2,*
1Division of Medical Oncology, Department of Internal Medicine, The Ohio State University, Columbus
2Division of Bioinformatics, Department of Biomedical Informatics, The Ohio State University, Columbus, USA
Ann Oncol. 2016 Mar; 27(3): 539–543.
Published online 2015 Dec 8.
Predicting Doxorubicin Response to Liposarcoma Using a Novel Genomic Matching Algorithm
Timothy Makkar, James L. Chen, MD
Department of Biomedical Informatics, The Ohio State University Wexner Medical Center
Opportunities for Patient Matching Algorithms to Improve Patient Care in Oncology
Travis Johnson, David Liebner, and James L. Chen
All authors: The Ohio State University, Columbus, OH
JCO Clinical Cancer Informatics 2017 :1, 1-8
Automating predictions of cancer patient outcomes
Jeffrey Spitzner, PhD1; Trevor Allen, BA1; Guy Jacks, BA1;James L Chen, MD1,2
1MatchTX, LLC, Columbus, Ohio
2Dept of Biomedical Informatics and Internal Medicine, The Ohio State University, Columbus, Ohio
CLICK HERE TO DOWNLOAD
a Case Study using MatchTx to predict outcome in High Gleason Score Prostate Cancer Patients.
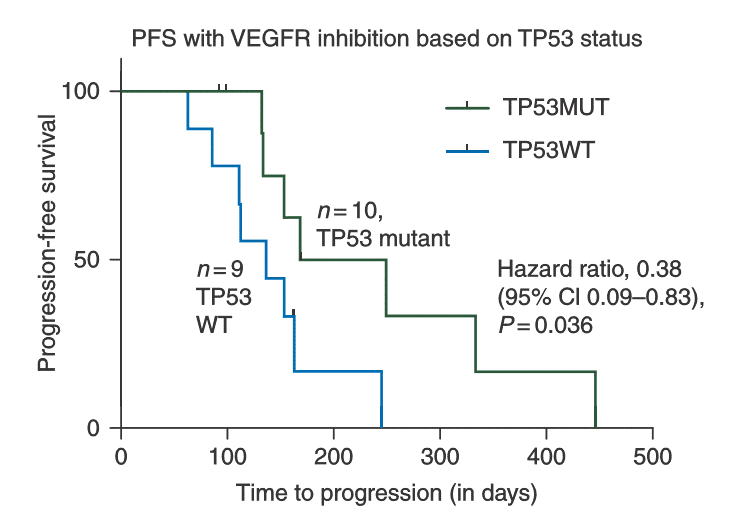
The MatchTx algorithms are able to predict the response of patients with advanced sarcomas to cancer drug pazopanib (a VEGFR inhibitor) based on the set of TP53 genetic mutations in the patient population. Taken from:
TP53 mutational status is predictive of pazopanib response in advanced sarcomas.
Click here to download a Poster presentation on the use of the MatchTx Technology
